Robert Langefeld: pyGEMMA: fast, scalable, user-friendly Python/R implementation of LMMs for GWAS скачать в хорошем качестве
Повторяем попытку...
Скачать видео с ютуб по ссылке или смотреть без блокировок на сайте: Robert Langefeld: pyGEMMA: fast, scalable, user-friendly Python/R implementation of LMMs for GWAS в качестве 4k
У нас вы можете посмотреть бесплатно Robert Langefeld: pyGEMMA: fast, scalable, user-friendly Python/R implementation of LMMs for GWAS или скачать в максимальном доступном качестве, видео которое было загружено на ютуб. Для загрузки выберите вариант из формы ниже:
-
Информация по загрузке:
Скачать mp3 с ютуба отдельным файлом. Бесплатный рингтон Robert Langefeld: pyGEMMA: fast, scalable, user-friendly Python/R implementation of LMMs for GWAS в формате MP3:
Если кнопки скачивания не
загрузились
НАЖМИТЕ ЗДЕСЬ или обновите страницу
Если возникают проблемы со скачиванием видео, пожалуйста напишите в поддержку по адресу внизу
страницы.
Спасибо за использование сервиса ClipSaver.ru
Robert Langefeld: pyGEMMA: fast, scalable, user-friendly Python/R implementation of LMMs for GWAS
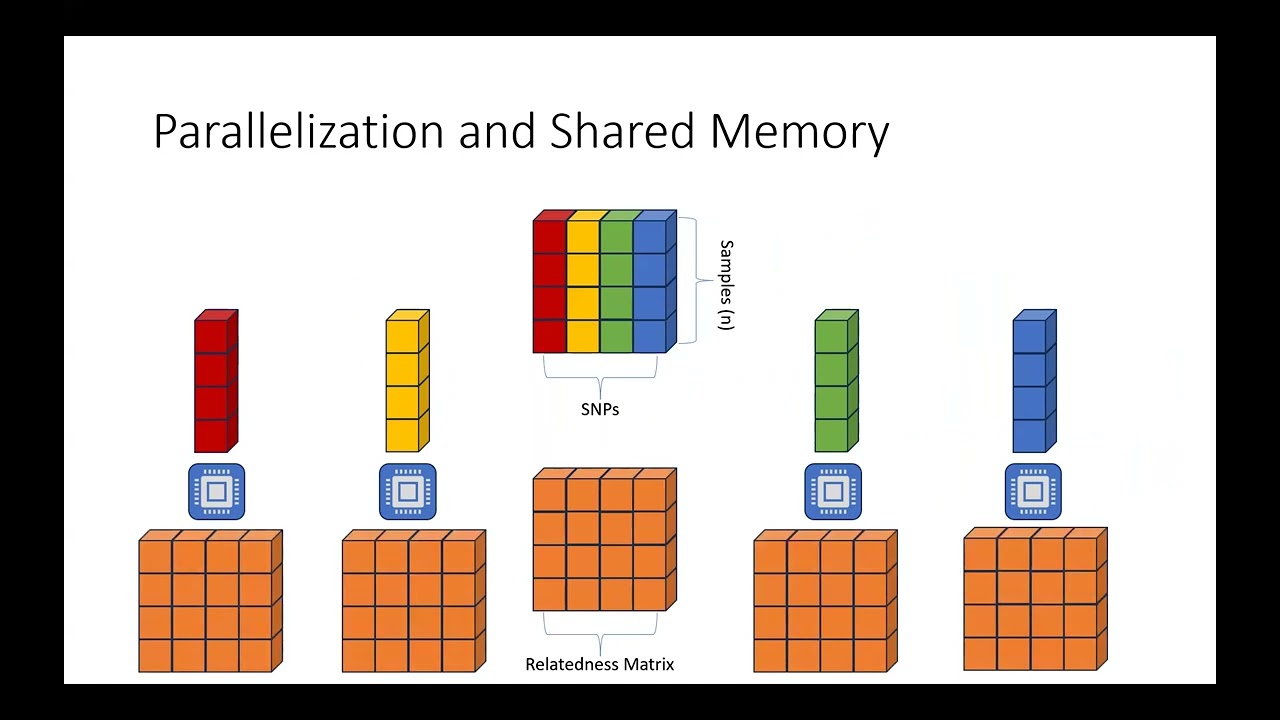
Abstract: Linear mixed models (LMMs) are widely used association analysis tools for genome-wide association studies (GWAS) to account for population stratification and relatedness among individuals. While multiple efficient LMM software packages like GEMMA, Bolt-LMM, and REGENIE exist, they often lack an accessible user interface, require specific data formats, and provide minimal parallel scaling. Here, we introduce pyGEMMA, a Python/Cython-based implementation of GEMMA for Python and R that provides a user-friendly framework for conducting LMM-based analyses in genetic studies with parallel computing capabilities. With pyGEMMA, researchers can perform end-to-end experiments with LMMs using high-level functions and scripts. pyGEMMA provides functions for both Python and R (through reticulate) for model-fitting and visualization without the specific data format requirements of other standard software. It includes functions for standard GWAS visualization (e.g., Manhattan plot, Q-Q plot). We also include computational efficiency improvements by leveraging tensor-based storage in NumPy to take advantage of additional BLAS and LAPACK operations. The package includes functionality to run numerous SNPs in parallel in order to scale to the hardware available. Additionally, we implement multi-node, scalable relatedness matrix calculation and eigendecomposition through the MPI communications protocol and SLATE (Software for Linear Algebra Targeting Exascale) library. We demonstrate the effectiveness of pyGEMMA through GWAS analyses on three datasets. We also demonstrate the scalability of pyGEMMA by running on large datasets of individuals of European descent from the UK Biobank. Our results show that pyGEMMA produces identical results to GEMMA with runtimes faster than competing methods while being more convenient and user-friendly. Through its multi-node computing approach, pyGEMMA's implementation also scales to larger datasets than most competing methods and maintains a reasonable runtime. By integrating GEMMA into Python and R, researchers can leverage its extensive functionality for LMM analysis while enjoying the ease and flexibility of Python/R programming. pyGEMMA is freely available on GitHub at github.com/rlangefe/pygemma. Bio: Robert Langefeld is a Ph.D. student in the University of Michigan Department of Biostatistics, working under Dr. Xiang Zhou. His field is statistical genetics and high performance computing.



















