Nickellaus Roberts-Sequencing of Multiple Displacement Amplification Products for Contiguous Genomes скачать в хорошем качестве
Повторяем попытку...

Скачать видео с ютуб по ссылке или смотреть без блокировок на сайте: Nickellaus Roberts-Sequencing of Multiple Displacement Amplification Products for Contiguous Genomes в качестве 4k
У нас вы можете посмотреть бесплатно Nickellaus Roberts-Sequencing of Multiple Displacement Amplification Products for Contiguous Genomes или скачать в максимальном доступном качестве, видео которое было загружено на ютуб. Для загрузки выберите вариант из формы ниже:
-
Информация по загрузке:
Скачать mp3 с ютуба отдельным файлом. Бесплатный рингтон Nickellaus Roberts-Sequencing of Multiple Displacement Amplification Products for Contiguous Genomes в формате MP3:
Если кнопки скачивания не
загрузились
НАЖМИТЕ ЗДЕСЬ или обновите страницу
Если возникают проблемы со скачиванием видео, пожалуйста напишите в поддержку по адресу внизу
страницы.
Спасибо за использование сервиса ClipSaver.ru
Nickellaus Roberts-Sequencing of Multiple Displacement Amplification Products for Contiguous Genomes
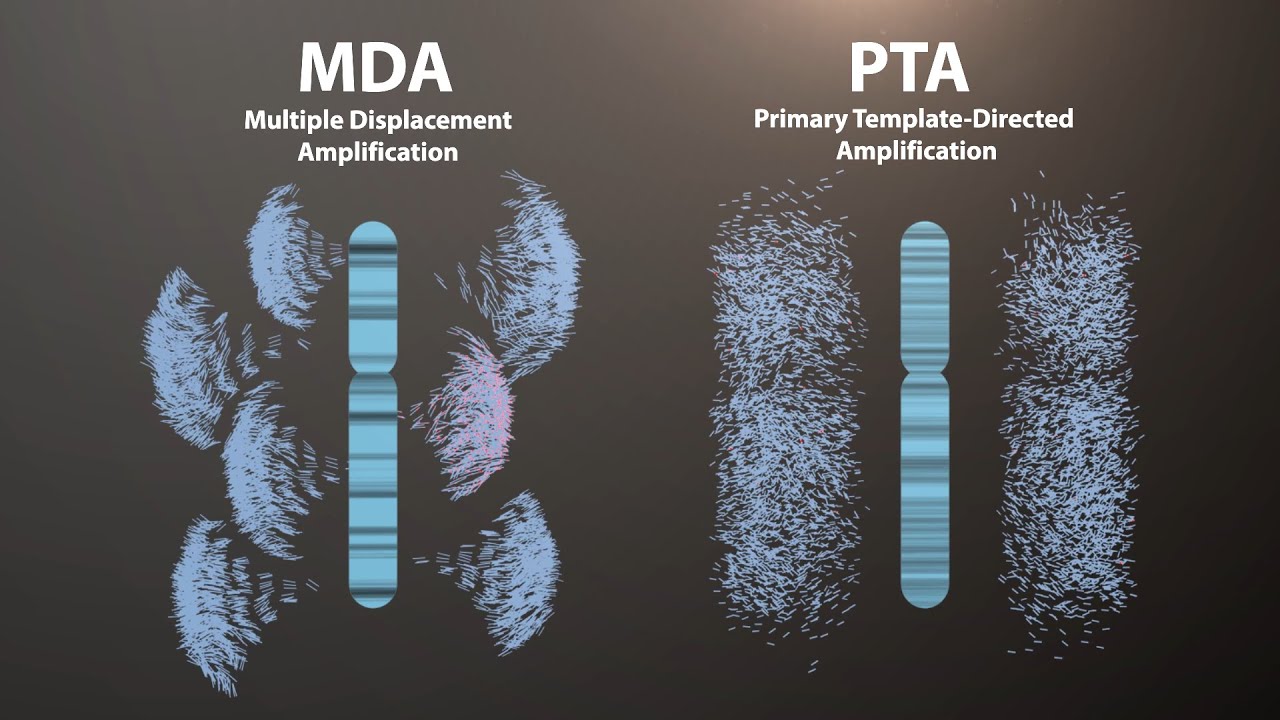
Advances in low-input sequencing library preparation for PacBio HiFi have enabled the sequencing of high-quality genomes from otherwise DNA-limited samples such as a single mosquito. However, in practice, even the Ultra-Low DNA Input Workflow for SMRT Sequencing requires more high-quality DNA than can be obtained from most meiofaunal invertebrates, which may be made up of fewer than 1,000 cells. Multiple displacement amplification (MDA) is the in vitro amplification of DNA using Phi29 DNA polymerase. Commercially available MDA kits can generate tens of micrograms of isothermally amplified DNA from a single meiofaunal animal. Attempts to sequence animal genomes from MDA DNA are sparse in the literature, but personal communications from several researchers who have attempted this indicate that Phi29’s bias in amplifying AT rich genomic regions and chimeras have previously posed major challenges. We sought to develop a workflow to solve these issues. Using a new, commercially available MDA kit marketed for single cell genomics and 1/2 PacBio HiFi cell per species, we sequenced highly contiguous genomes of two individual microscopic invertebrates. The method was assessed using the model organism Caenorhabditis elegans. The Caenorhabditis elegans assembly contains 336 contigs (N50 = 868 Kbp, L50 = 39) and a BUSCO score of 94.9% (nematoda_odb10). Furthermore, a non-reference guided Caenorhabditis elegans assembly when mapped to the chromosomes of the reference resulted in an average 94.68% coverage per chromosome. Read mapping also indicates very low rates of chimeric reads. The de novo assembly of Lepidodermella squamata tested with this method resulted in an assembly containing 527 contigs (N50 = 2.9 Mbp, L50 = 17) and a BUSCO score of 80.1% (metazoa_odb10). These results demonstrate that MDA may be an easy and cost-effective way to produce high quality genomes from microscopic animals with small genomes although more tests on taxa with larger genomes are needed.