Inferring bacterial cell biology and gene regulation from transcriptional "noise" скачать в хорошем качестве
Повторяем попытку...

Скачать видео с ютуб по ссылке или смотреть без блокировок на сайте: Inferring bacterial cell biology and gene regulation from transcriptional "noise" в качестве 4k
У нас вы можете посмотреть бесплатно Inferring bacterial cell biology and gene regulation from transcriptional "noise" или скачать в максимальном доступном качестве, видео которое было загружено на ютуб. Для загрузки выберите вариант из формы ниже:
-
Информация по загрузке:
Скачать mp3 с ютуба отдельным файлом. Бесплатный рингтон Inferring bacterial cell biology and gene regulation from transcriptional "noise" в формате MP3:
Если кнопки скачивания не
загрузились
НАЖМИТЕ ЗДЕСЬ или обновите страницу
Если возникают проблемы со скачиванием видео, пожалуйста напишите в поддержку по адресу внизу
страницы.
Спасибо за использование сервиса ClipSaver.ru
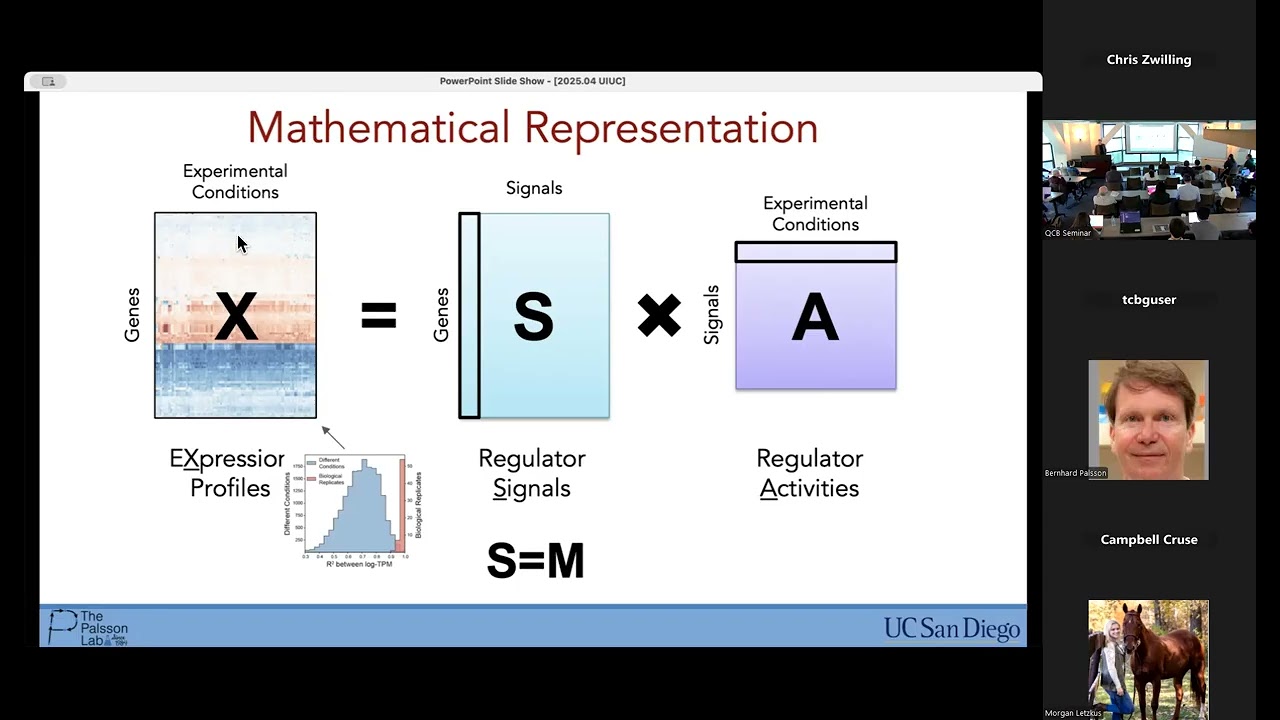
Inferring bacterial cell biology and gene regulation from transcriptional "noise"
QCB Seminar April 18 2025. Andrew Pountain, UT Health, Houston. Gene expression varies between individual cells, even those with identical genomes in a homogeneous environment. In bacteria, our understanding of what drives this variation has been obscured by a lack of methods to measure single-cell gene expression at the genome-wide scale. By applying a recently developed single cell RNA sequencing protocol, we showed that interactions between transcription and chromosomal replication drive global heterogeneity in gene expression. Analyzing genome-wide patterns of covariance, we could reconstruct DNA replication patterns and use these to infer cell cycle transcriptional dynamics in these organisms for the first time. We found that replication itself impacts transcription of each gene, but that the response of individual genes to this perturbation is shaped by a gene’s local regulatory context, from its turnover rate to its repression state. We are continuing to develop our understanding of transcription-replication interactions, as well as expanding our analysis across different states of growth. Overall, bacterial transcriptional variation, far from being simply “noise”, contains quantitatively reproducible information about the state of the underlying biological system, far beyond what can be inferred from bulk measurement.